A team of scientists from Michal Hocek Group at IOCB Prague, Miroslav Fojta Group at the Institute of Biophysics of the Czech Academy of Sciences in Brno and Ciara O’Sullivan from the Universitat Rovira i Vigili in Tarragona have developed an improved method for the redox labeling of nucleotides in DNA for electrochemical detection.
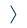
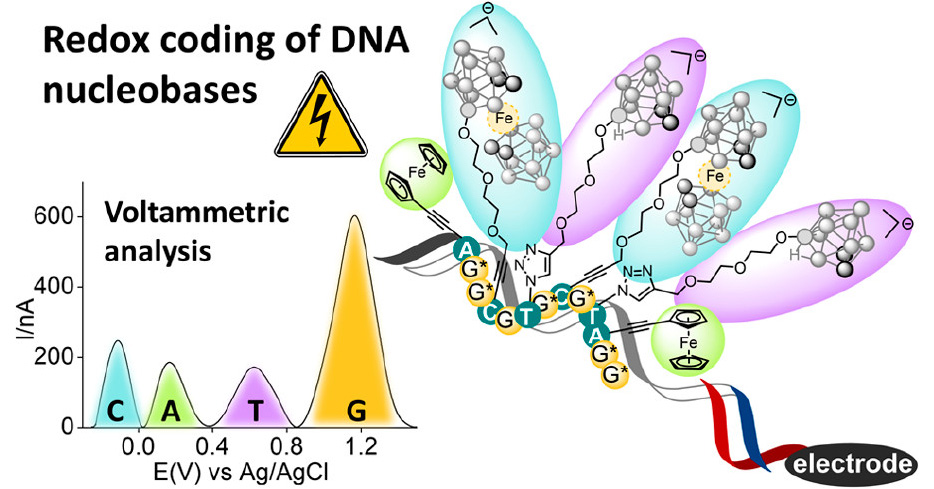
They used differently substituted ferrocene derivatives to tune their redox potential, making the labeling moieties behave orthogonally. Therefore, they enabled the quantification of the nucleobases derivatized in this way via the square-wave voltammetry.
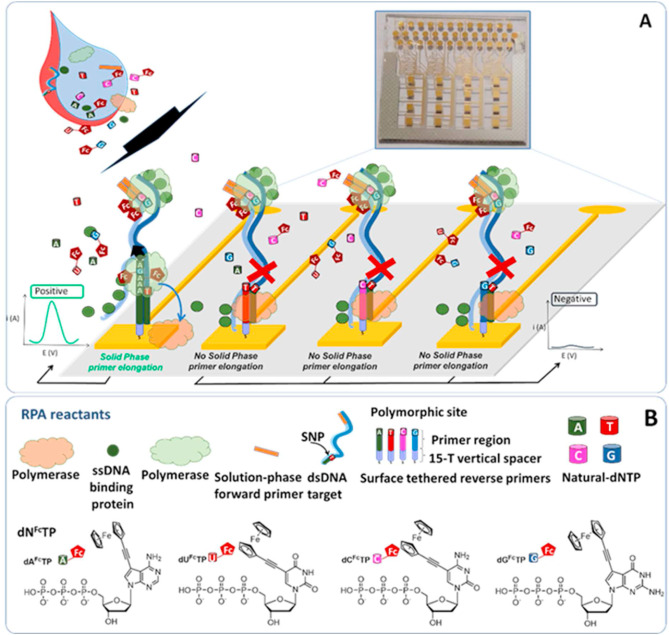
Scientists designed and synthesized three ferrocene-based labels, a plain ferrocene moiety, ferrocenecarboxamide and octamethylferrocene, all of them bound to the nucleobase through the ethynyl linker. The modified nucleotides were incorporated into a DNA segment using the KOD XL DNA polymerase and the labeled DNA was analyzed on the gold electrode using capture probes.
The octamethylferrocene group was found to be unsuitable for quantification because it is prone to oxidation by the air, but the ferrocene and ferrocenecarboxamide worked well and were ratiometrically detected in the presence of the other label.
This chemical tool could find its application as a label to directly measure the relative ratio of the nucleobases in an unknown sequence of the DNA, using orthogonal ferrocene labels with different redox potentials.
Original paper
- Simonova, A., Magriñá, I., Sýkorová, V., Pohl, R., Ortiz, M., Havran, L., Fojta, M., O'Sullivan, C.K. and Hocek, M. (2020), Tuning of Oxidation Potential of Ferrocene for Ratiometric Redox Labeling and Coding of Nucleotides and DNA. Chem. Eur. J. https://doi.org/10.1002/chem.201904700
This research was co-funded by OP RDE project ChemBioDrug No. CZ.02.1.01/0.0/0.0/16_019/0000729.