Saccharides have multiple significant roles in living organisms. The most known are energy storage, mechanical support, and as a scaffold for genetic information. They also participate in cell-cell recognition and signaling, often in combination with proteins. To fully grasp these roles, we need to determine their 3D structures and understand their conformational behavior in water solutions. Saccharides, however, are notorious for being difficult to study using traditional structural biology approaches, such as NMR, UV-vis spectroscopy, and X-ray scattering techniques.
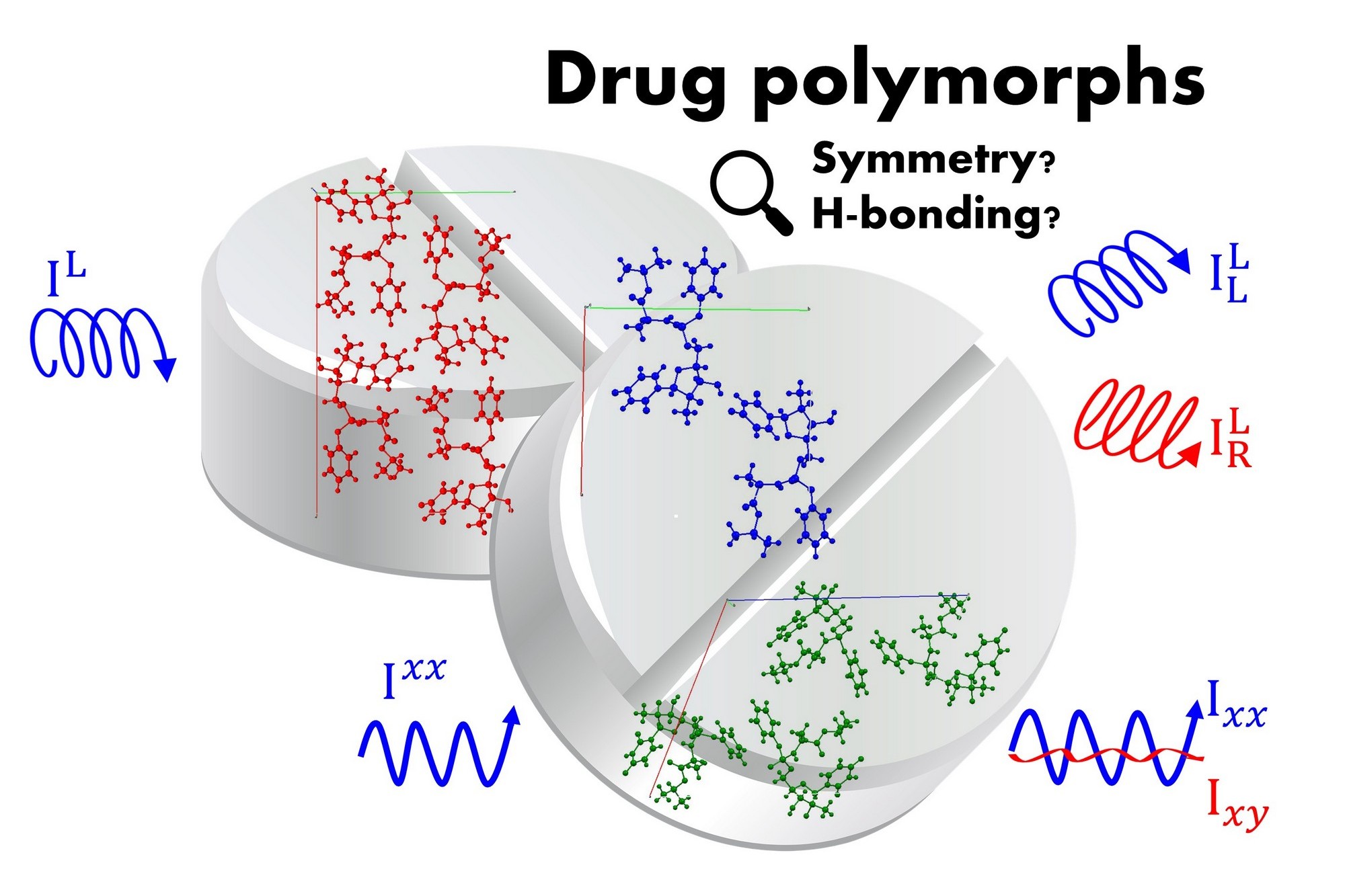
Raman and Raman optical activity spectroscopies were proposed as suitable techniques sensitive to the structural features of saccharides. The problem is that the resulting experimental spectra for both techniques are insufficient to decipher the desired structural molecular information. In fact, these methods greatly benefit from computational modeling used for predictions of the spectra. Such predictions, however, need to consider the flexibility of the saccharide molecule as well as its interaction with nearby water molecules, which makes these calculations extremely complicated.
Vladimír Palivec, Jakub Kaminský, and Hector Martinez-Seara, together with other colleagues, developed a transferable protocol for such calculations that provides accurate predictions and, unlike its predecessors, takes up significantly fewer computational resources. This means the protocol can face even larger oligosaccharides without sacrificing accuracy.
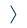
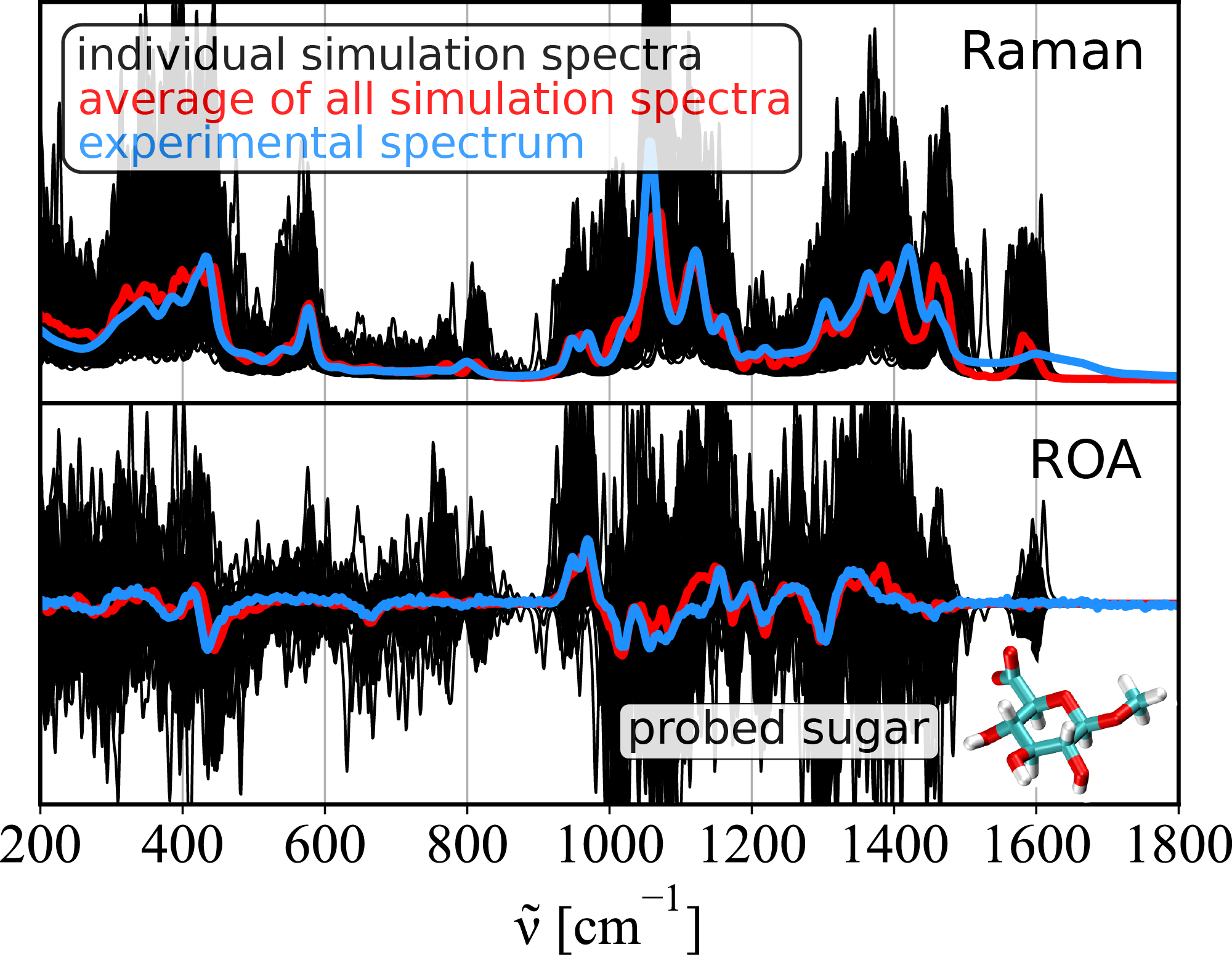
The key idea of the proposed approach is to account for the real variability of the structures of the sugars and their water surroundings in the spectra calculation. First, it uses fast molecular dynamics simulations to gather multiple conformational snapshots. This is a cheap approach based on a classical description of the molecules (i.e. balls and springs). Then, for each selected structure, the spectrum is obtained using the quantum chemistry computation. The experimental spectra prediction then comes as an average of the computed spectra weighted by the relative conformation abundance.
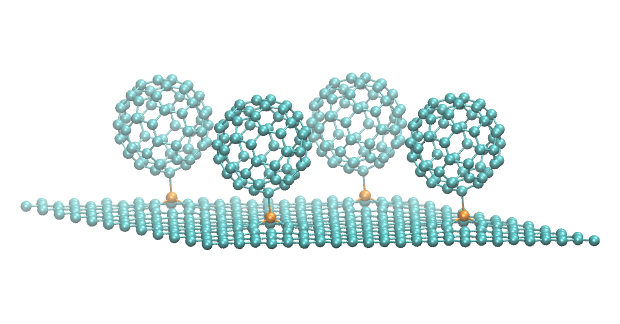
Moreover, the approach employs a hybrid quantum mechanics/molecular mechanics method, improving both its computational efficiency and accuracy. The advantage is that only the central saccharide requires the costly quantum description, while the explicitly considered water molecules found around the sugar use a significantly cheaper molecular mechanics description.
The proposed methodology corrects imperfections connected with the applied quantum theory methods and their need for scaling calculated vibrational frequencies, allowing for an excellent match of simulation on experimental data.
Compared to the experimental data, the simulation gives highly accurate spectral overlap and captures most of the spectral features overall. Furthermore, the method is aware of the underlying 3D-structures and provides full structural determination, making it a potentially useful tool for the study of saccharides in their biologically active environments.