Researchers at IOCB Prague develop a new method for enzymatic synthesis of potential RNA therapeutics

A team of researchers at IOCB Prague led by Prof. Michal Hocek has developed a novel method for preparing ribonucleic acid (RNA) containing modified bases. Innovative use of engineered DNA polymerases, enzymes commonly used for synthesis of DNA, led to development of a general method for the synthesis of RNA modified only at selected sites or even RNA containing various modifications at all nucleotide building blocks. This opens the door to applications in chemical biology and, in a longer perspective, in the therapy of hitherto incurable diseases. The paper was published in the journal Nature Communications.
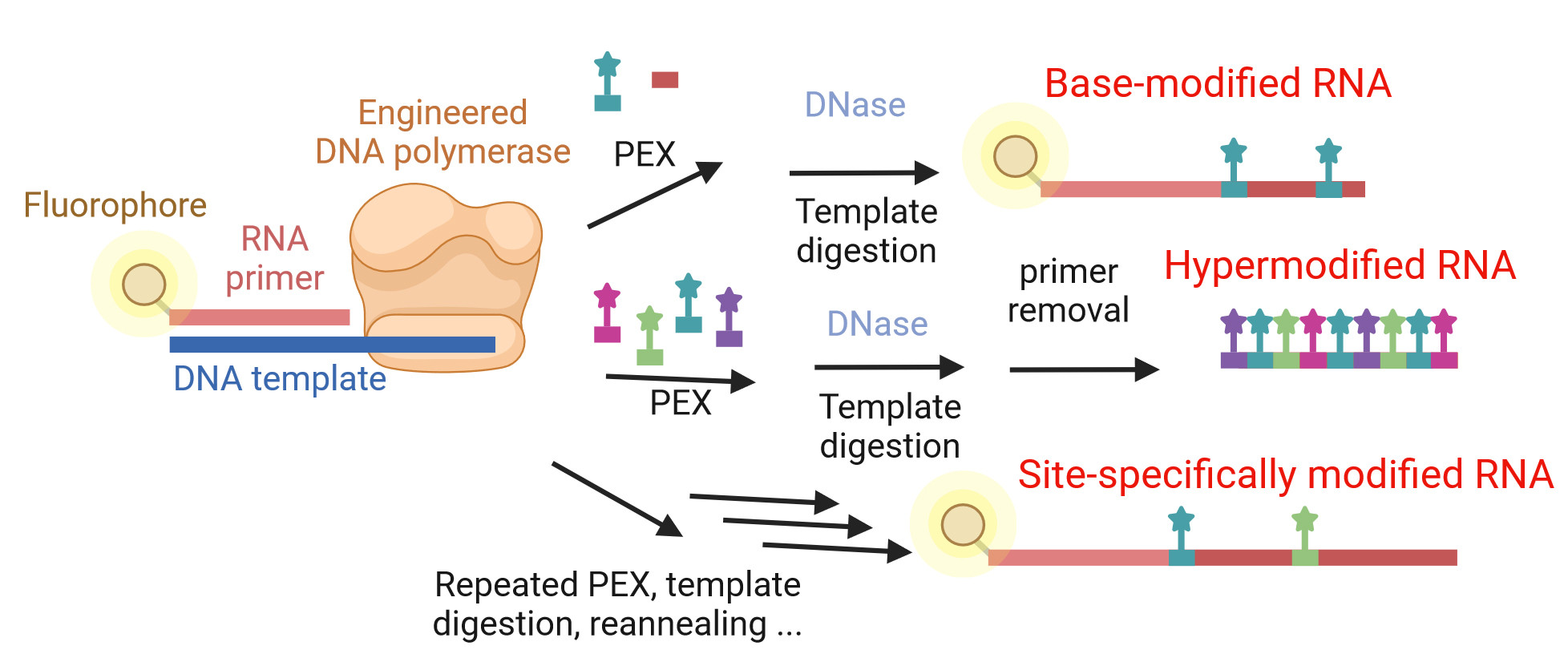
The researchers used two artificially modified DNA polymerases known to be capable of synthesizing RNA. In nature, DNA polymerases synthesize only DNA, while RNA polymerases produce RNA. Hocek’s team has developed a procedure for preparing modified RNA with significant advantages over the commonly available method, in vitro transcription with viral T7 RNA polymerase, which, for example, is used in the production of well-known mRNA vaccines.
Modified DNA polymerases can incorporate virtually any modification into any RNA sequence, even only at selected specific sites, precisely where the given modification is required. The commonly used T7 RNA polymerase can only incorporate the modified base in all positions in the RNA but not in one precisely designated location. Therefore, if the RNA requires site-specific modification, the classical in vitro transcription fails and a different method is needed.
And that’s exactly what the team at IOCB has developed, a method that in many respects is comparable to the commonly used in vitro transcription while eliminating most of its pitfalls and offering completely new possibilities. Now, various specific RNA probes can be prepared for the study of RNA biology, which is currently a very hot topic. In the longer term, however, there is great promise for therapeutic applications, especially for mRNA therapeutics.
The researchers chose two specific positions in mRNA for modifications and found that they led to a significant increase in the production of certain proteins. This is an excellent news for the development of potential mRNA drugs. If mRNA modified in this way could be introduced into cells, it would be possible to trigger the production of a protein which the body lacks or which is defective.
“Our method may lead to the development of therapeutics for the treatment of many diseases, including cancer and some genetic diseases caused by a malfunctioning or missing protein. It allows for the replacement of a missing or poorly functioning protein,” says Michal Hocek. He continues: “RNA therapy is a powerful technology, and it may emerge as one of the main directions of drug development within the next ten years.”
The challenge is how to ensure that there is just the right amount of protein – not scarce and neither excessive – at just the right time. In most cases, if there is too much of certain proteins, they are harmful to the body. That’s why mRNA naturally breaks down quickly in the cell and the entire process is very finely regulated by the body. According to Michal Hocek, their method will not replace the classical in vitro transcription. However, if there is a need for specific modifications to occur only at selected RNA sites, then the advantage will lie with the novel method from IOCB Prague.
Original article: Brunderová, M.; Havlíček, V.; Matyašovský, J.; Pohl, R.; Poštová Slavětínská, L.; Krömer, M.; Hocek, M. Expedient production of site specifically nucleobase-labelled or hypermodified RNA with engineered thermophilic DNA polymerases. Nat. Commun. 2024, 15, 3054. https://doi.org/10.1038/s41467-024-47444-9